Tutorial - Module 2: Accessibility Analysis#
This notebook is a template workflow to collect data and prepare the main data to perform a baseline physical accessibility analysis to health facilities. It uses various tools developed by the World Bank’s Geospatial Operations Support Team (GOST).
This notebook represents the second module of the tutorial on the physical accessibility to health facilities in a context of emergency.
It focuses on the datasets and the functions to be used to perform the analysis. A particular focus will be put on the uncertainty and pros&cons deriving from the utilization of different datasets. \
Setup#
Import packages required for the analysis
# System
import sys
import os
from os.path import join, expanduser
from pathlib import Path
# Avoid warnings to pop up
import warnings
warnings.filterwarnings('ignore')
# Visualization tools
# import folium as flm
import matplotlib.pyplot as plt
import matplotlib.colors as colors
import matplotlib.gridspec as gridspec
from rasterio.plot import plotting_extent
from rasterio.plot import show
from mpl_toolkits.axes_grid1 import make_axes_locatable
import contextily as ctx
import cartopy
import cartopy.crs as ccrs
import cartopy.feature as cfeature
import seaborn as sns
os.environ['CARTOPY_USER_BACKGROUNDS'] = '/home/jupyter-wb618081/Python/Backgrounds/'
# Processing
import numpy as np
import geopandas as gpd
import pandas as pd
from gadm import GADMDownloader
import dask_geopandas as dask_gpd
# Raster
import rasterio as rio
from rasterio.features import shapes
from shapely.geometry import box
from rasterio.features import geometry_mask
from rasterstats import zonal_stats
from shapely.geometry import Polygon, box, Point
from shapely.geometry import mapping
import skimage.graph as graph
from scipy.signal import convolve2d
# Graph
import pickle
import networkx as nx
import osmnx as ox
# for facebook data
from pyquadkey2 import quadkey
# Climate/Flood
# import xarray as xr
# Define your path to the Repositories
sys.path.append(join(expanduser("/home/jupyter-wb618081"), 'Repos', 'gostrocks', 'src'))
sys.path.append(join(expanduser("/home/jupyter-wb618081"), 'Repos', 'GOSTNets_Raster', 'src'))
sys.path.append(join(expanduser("/home/jupyter-wb618081"), 'Repos', 'GOSTnets'))
sys.path.append(join(expanduser("/home/jupyter-wb618081"), 'Repos', 'GOST_Urban', 'src', 'GOST_Urban'))
sys.path.append(join(expanduser("/home/jupyter-wb618081"), 'Repos', 'health-equity-diagnostics', 'src', 'modules'))
sys.path.append(join(expanduser("/home/jupyter-wb618081"), 'Repos', 'INFRA_SAP'))
import GOSTnets as gn
from GOSTnets.load_osm import *
import GOSTRocks.rasterMisc as rMisc
from GOSTRocks.misc import get_utm
import GOSTNetsRaster.market_access as ma
import UrbanRaster as urban
from infrasap import aggregator
from infrasap import osm_extractor as osm
from utils import download_osm_shapefiles
# auto reload
%load_ext autoreload
%autoreload 2
Define below the local folder where you are located
scratch_dir = join(expanduser("/home/jupyter-wb618081"), 'Health-Access-Metrics', 'Tutorials')
data_dir = join(scratch_dir, 'tutorial_data')
out_path = join(scratch_dir, 'tutorial_output')
## Function for creating a path, if needed ##
def checkDir(out_path):
if not os.path.exists(out_path):
os.makedirs(out_path)
Data Preparation#
Import the City boundaries (.shp)#
epsg = "EPSG:4326"
epsg_utm = "EPSG:28232"
country = 'Congo'
iso = 'COG'
city = gpd.read_file((data_dir+"/brazaville.shp"))
city.to_crs(epsg)
city
| ID_0 | COUNTRY | NAME_1 | NL_NAME_1 | ID_2 | NAME_2 | VARNAME_2 | NL_NAME_2 | TYPE_2 | ENGTYPE_2 | CC_2 | HASC_2 | geometry | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | COG | Republic of the Congo | Brazzaville | NaN | COG.2.1_1 | Brazzaville | NaN | NaN | District | District | NaN | CG.BR.BR | POLYGON ((15.32458 -4.28091, 15.31317 -4.27887... |
Import the Road Network (.shp)#
Download from the link above the OpenStreetMap road network from Geofabrik
roads_osm = OSM_to_network(join(data_dir,"congo-brazzaville-latest.osm.pbf"))
GOSTNets creates a special ‘OSM_to_network’ object. This object gets initialized with both a copy of the OSM file itself and the roads extracted from the OSM file in a GeoPandas DataFrame. This DataFrame is a property of the object called ‘roads_raw’ and is the starting point for our network.
?roads_osm
Type: OSM_to_network
String form: <GOSTnets.load_osm.OSM_to_network object at 0x7ec141828b20>
File: ~/Repos/GOSTnets/GOSTnets/load_osm.py
Docstring:
Object to load OSM PBF to networkX objects.
Object to load OSM PBF to networkX objects. EXAMPLE: G_loader = losm.OSM_to_network(bufferedOSM_pbf) G_loader.generateRoadsGDF() G = G.initialReadIn()
snap origins and destinations o_snapped = gn.pandana_snap(G, origins) d_snapped = gn.pandana_snap(G, destinations)
Init docstring: Generate a networkX object from a osm file
roads_gdf = roads_osm.roads_raw
roads_osm.roads_raw.head()
| osm_id | infra_type | one_way | bridge | geometry | |
|---|---|---|---|---|---|
| 0 | 4692113 | primary | True | False | LINESTRING (15.23126 -4.29689, 15.23134 -4.296... |
| 1 | 4692179 | primary | True | False | LINESTRING (15.25202 -4.28510, 15.25210 -4.285... |
| 2 | 4692181 | primary | True | False | LINESTRING (15.24010 -4.29174, 15.23988 -4.291... |
| 3 | 4692231 | primary | True | False | LINESTRING (15.28165 -4.27427, 15.28176 -4.274... |
| 4 | 4692245 | primary | True | False | LINESTRING (15.25944 -4.28030, 15.25946 -4.280... |
?roads_osm.roads_raw
Type: GeoDataFrame
String form:
osm_id infra_type one_way bridge \
0 4692113 primary True Fals <...> -4.785...
54721 LINESTRING (11.94277 -4.78254, 11.94306 -4.784...
[54722 rows x 5 columns]
Length: 54722
File: ~/.conda/envs/geo_wb_linux/lib/python3.8/site-packages/geopandas/geodataframe.py
Docstring:
A GeoDataFrame object is a pandas.DataFrame that has a column
with geometry. In addition to the standard DataFrame constructor arguments,
GeoDataFrame also accepts the following keyword arguments:
Parameters
----------
crs : value (optional)
Coordinate Reference System of the geometry objects. Can be anything accepted by
:meth:`pyproj.CRS.from_user_input() <pyproj.crs.CRS.from_user_input>`,
such as an authority string (eg "EPSG:4326") or a WKT string.
geometry : str or array (optional)
If str, column to use as geometry. If array, will be set as 'geometry'
column on GeoDataFrame.
Examples
--------
Constructing GeoDataFrame from a dictionary.
>>> from shapely.geometry import Point
>>> d = {'col1': ['name1', 'name2'], 'geometry': [Point(1, 2), Point(2, 1)]}
>>> gdf = geopandas.GeoDataFrame(d, crs="EPSG:4326")
>>> gdf
col1 geometry
0 name1 POINT (1.00000 2.00000)
1 name2 POINT (2.00000 1.00000)
Notice that the inferred dtype of 'geometry' columns is geometry.
>>> gdf.dtypes
col1 object
geometry geometry
dtype: object
Constructing GeoDataFrame from a pandas DataFrame with a column of WKT geometries:
>>> import pandas as pd
>>> d = {'col1': ['name1', 'name2'], 'wkt': ['POINT (1 2)', 'POINT (2 1)']}
>>> df = pd.DataFrame(d)
>>> gs = geopandas.GeoSeries.from_wkt(df['wkt'])
>>> gdf = geopandas.GeoDataFrame(df, geometry=gs, crs="EPSG:4326")
>>> gdf
col1 wkt geometry
0 name1 POINT (1 2) POINT (1.00000 2.00000)
1 name2 POINT (2 1) POINT (2.00000 1.00000)
See also
--------
GeoSeries : Series object designed to store shapely geometry objects
roads_osm.roads_raw.infra_type.value_counts()
infra_type
residential 35414
track 7667
unclassified 3599
path 2504
service 1696
tertiary 1608
trunk 746
secondary 655
primary 409
footway 297
primary_link 36
trunk_link 27
construction 21
tertiary_link 20
secondary_link 9
living_street 5
steps 4
pedestrian 3
yes 2
Name: count, dtype: int64
We can show the different highway types and counts
We need to define a list of the types of roads from the above that we consider acceptable for our road network
accepted_road_types = ['residential', 'track','unclassified','path', 'service','tertiary','trunk','secondary','primary',
'footway','primary_link','trunk_link','secondary_link','tertiary_link']
We can therefore filter our roads using the filterRoads method
roads_osm.filterRoads(acceptedRoads = accepted_road_types)
roads_osm.roads_raw.infra_type.value_counts()
infra_type
residential 35414
track 7667
unclassified 3599
path 2504
service 1696
tertiary 1608
trunk 746
secondary 655
primary 409
footway 297
primary_link 36
trunk_link 27
tertiary_link 20
secondary_link 9
Name: count, dtype: int64
We are interested in the area of the capital city of Congo, Brazzaville. We therefore clip the road network to the shapefile of the city
# This is the GeoPandas dataframe
display(city)
city.plot()
| ID_0 | COUNTRY | NAME_1 | NL_NAME_1 | ID_2 | NAME_2 | VARNAME_2 | NL_NAME_2 | TYPE_2 | ENGTYPE_2 | CC_2 | HASC_2 | geometry | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | COG | Republic of the Congo | Brazzaville | NaN | COG.2.1_1 | Brazzaville | NaN | NaN | District | District | NaN | CG.BR.BR | POLYGON ((15.32458 -4.28091, 15.31317 -4.27887... |
<Axes: >
# This is the Shapely geometry object contained in the geodf
city_shp = city.geometry[0]
city_shp
We check to see everything lines up by running intersect and counting the True / False returns. The count of the True values are the number of roads that intersect the AOI
intersects is a Shapely function that returns True if the boundary or interior of the object intersect in any way with those of the other
roads_osm.roads_raw.to_crs(epsg)
print(roads_osm.roads_raw.crs)
print(city.crs)
+init=epsg:4326 +type=crs
EPSG:4326
roads_osm.roads_raw.geometry.intersects(city_shp).value_counts()
False 48701
True 5986
Name: count, dtype: int64
We can therefore remove any roads that does not intersect Brazzaville administrative unit
roads_osm.roads_raw = roads_osm.roads_raw.loc[roads_osm.roads_raw.geometry.intersects(city_shp) == True]
Now we generate the RoadsGPD object, which is stored as a property of the ‘OSM_to_network’ object. The RoadsGPD object is a GeoDataFrame that further processes the roads.
This includes splitting the edges where intersections occur, adding unique edge IDs, and adding to/from columns to the GeoDataFrame.
We can do this using the generateRoadsGDF function
roads_osm.generateRoadsGDF(verbose = False)
We use the initialReadIn() function to transform this to a graph object
roads_osm.initialReadIn()
<networkx.classes.multidigraph.MultiDiGraph at 0x7ebd20adefd0>
roads_osm.network
<networkx.classes.multidigraph.MultiDiGraph at 0x7ebd20adefd0>
We save this graph object down to file using gn.save().
The save function produces three outputs: a node GeoDataFrame as a CSV, an edge GeoDataFrame as a CSV, and a graph object saved as a pickle (check your folder!).
gn.save(roads_osm.network,'roads_brazzaville',out_path)
Import Friction Surface (.tif)#
travel_surf = rio.open(join(data_dir, f"travel_surface_motorized_2020_{iso}.tif"))
travel_surf.res
(0.008333333333333333, 0.008333333333333333)
Import the Health Facilities (destinations)#
Health facilities are stored as Geopandas dataframe
hf = gpd.read_file((data_dir+"/hf_COG.shp"))
hf = hf[hf.geometry.intersects(city_shp)]
display('The following categories and numbers of Health Facilities are considered to perform the analysis: ')
display(hf["Facility t"].value_counts())
'The following categories and numbers of Health Facilities are considered to perform the analysis: '
Facility t
Centre de Santé Intégré 22
l?Hôpital de Base 2
University Hospital 1
Name: count, dtype: int64
Import Population (origin)#
# wp_path = join(expanduser("R:/"), 'Data', 'GLOBAL/Population/WorldPop_PPP_2020/MOSAIC_ppp_prj_2020', f'ppp_prj_2020_{iso}.tif') # Download from link above
wp_path = join(data_dir, f'cog_ppp_2020_1km_Aggregated.tif') # Download from link above
pop_surf = rio.open(wp_path)
rMisc.clipRaster(pop_surf, city, join(data_dir, f"pop_brazaville.tif"), crop=True)
wp_path = join(data_dir, f'pop_brazaville.tif')
pop_surf = rio.open(wp_path)
# Create a population df from population surface
indices = list(np.ndindex(pop_surf.shape))
xys = [Point(pop_surf.xy(ind[0], ind[1])) for ind in indices]
res_df = pd.DataFrame({
'spatial_index': indices,
'geometry': xys,
'pop': pop_surf.read(1).flatten()
})
res_df = res_df[res_df["pop"] != -99999.0]
res_df.head(2)
| spatial_index | geometry | pop | |
|---|---|---|---|
| 8 | (0, 8) | POINT (15.277916606793564 -4.204583108029534) | 7612.103027 |
| 9 | (0, 9) | POINT (15.286249940093564 -4.204583108029534) | 8549.582031 |
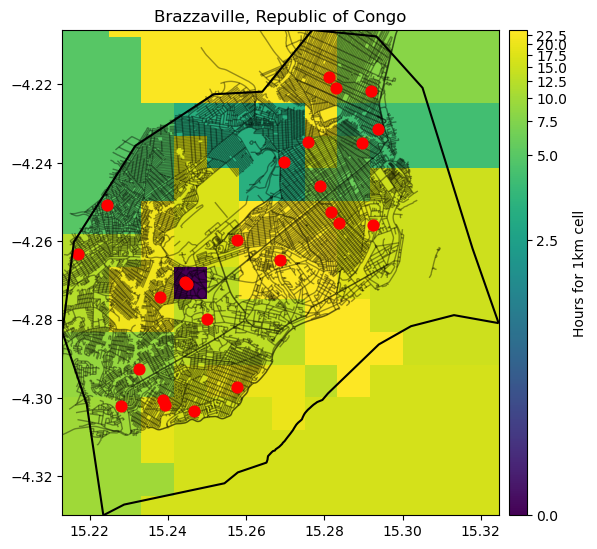
Let’s plot the location of the Health facilities, together with the travel surface dataset and the roads
fig, ax = plt.subplots(figsize=(6, 8))
ax.set_title("Brazzaville, Republic of Congo", fontsize=12, horizontalalignment='center')
# Plot the flood data
pop_image = show(marketsheds, transform=travel_surf.transform, ax=ax, norm=colors.PowerNorm(gamma=0.15), cmap='viridis', alpha = 1, zorder = 2)
# travel_surf.read(1)*1000/60
# Create an axis divider for the colorbar
divider = make_axes_locatable(ax)
cax = divider.append_axes('right', size="4%", pad=0.1)
# Add the colorbar
cb = fig.colorbar(pop_image.get_images()[0], cax=cax, orientation='vertical')
cb.set_label("Hours for 1km cell")
# Plot the road network not impcted
roads_osm.roads_raw.plot(ax=ax, color='black', linewidth=1, alpha = 0.4, zorder = 2)
# PLot the Health facilities
hf.plot(ax=ax, color='red', markersize = 60, zorder=3)
city.boundary.plot(ax=ax, color = "black", zorder=3, edgecolor="black")
ax.set_xlim(city_shp.bounds[0], city_shp.bounds[2])
ax.set_ylim(city_shp.bounds[1], city_shp.bounds[3])
plt.show()
Calculate Travel Time#
with open(os.path.join(out_path, 'roads_brazzaville.gpickle'), 'rb') as f:
G = pickle.load(f)
print('start: %s\n' % time.ctime())
G_clean = gn.clean_network(G, wpath = out_path, output_file_name = 'roads_brazzaville_clean', UTM = epsg_utm, WGS = epsg, junctdist = 10, verbose = True)
# using verbose = True:
# G_clean = gn.clean_network(G, wpath = data_pth, output_file_name = 'iceland_network', UTM = Iceland_UTMZ, WGS = {'init': 'epsg:4326'}, junctdist = 10, verbose = True)
print('\nend: %s' % time.ctime())
print('\n--- processing complete')
start: Thu Sep 12 10:56:30 2024
21554
completed processing 43108 edges
20985
completed processing 41970 edges
Edge reduction: 21554 to 41970 (-94 percent)
end: Thu Sep 12 10:56:57 2024
--- processing complete
Each edge in the network has a property called ‘length’. This was actually computed during Step 1 when the generateRoadsGDF function was run. The units of this length are in kilometres.
We can convert length to time, so that to calculate how long it takes to reach a destination, using the convert_network_to_time function.
We use a factor of 1000, because the function is expecting meters, so we need to convert the units of kilometers to meters.
The convert_network_to_time function uses a default speed dictionary that assigns speed limits to OSM highway types. However, it is possible to specify your own speed dictionary.
G_time = gn.convert_network_to_time(G_clean, distance_tag = 'length', road_col = 'infra_type', factor = 1000)
gn.example_edge(G_time)
(0, 1325, {'Wkt': 'LINESTRING (15.2475657 -4.2920382, 15.2467013 -4.2919278)', 'id': 2406, 'infra_type': 'residential', 'one_way': False, 'osm_id': '48569609', 'key': 'edge_2406', 'length': 96.72998683022037, 'Type': 'legitimate', 'time': 17.411397629439666, 'mode': 'drive'})
In order to perform network analysis we want to know the closest network node to each hospital.
For this, we use the pandana_snap_c function to snap (project and retrive closest node distance) the hospitals locations to the road network:
gn.pandana_snap_c?
Signature:
gn.pandana_snap_c(
G,
point_gdf,
source_crs='epsg:4326',
target_crs='epsg:4326',
add_dist_to_node_col=True,
time_it=False,
)
Docstring:
snaps points to a graph at a faster speed than pandana_snap.
:param G: a graph object, or the node geodataframe of a graph
:param point_gdf: a geodataframe of points, in the same source crs as the geometry of the graph object
:param source_crs: The crs for the input G and input point_gdf in format 'epsg:32638'
:param target_crs: The measure crs how distances between points are calculated. The returned point GeoDataFrame's CRS does not get modified. The crs object in format 'epsg:32638'
:param add_dist_to_node_col: return distance to nearest node in the units of the target_crs
:param time_it: return time to complete function
:return: returns a GeoDataFrame that is the same as the input point_gdf but adds a column containing the id of the nearest node in the graph, and the distance if add_dist_to_node_col == True
File: ~/Repos/GOSTnets/GOSTnets/core.py
Type: function
destinations = gn.pandana_snap_c(G_time, hf, source_crs = epsg, target_crs = epsg_utm)
destinations.head(2)
| Country | Admin1 | Facility n | Facility t | Ownership | Lat | Long | LL source | geometry | NN | NN_dist | |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 23 | Congo | Brazzaville | 3 Martyrs Centre de Santé Intégré | Centre de Santé Intégré | MoH | -4.2704 | 15.2443 | Combination | POINT (15.24430 -4.27040) | 3167 | 45.510237 |
| 24 | Congo | Brazzaville | Bissita Centre de Santé Intégré | Centre de Santé Intégré | MoH | -4.3005 | 15.2387 | Google Earth | POINT (15.23870 -4.30050) | 12530 | 58.768015 |
When calculating an OD-Matrix, we can only use the node IDs as inputs. So, we convert this column of our dataframe over to a list of unique values:
destinations = list(set(destinations.NN))
Now let’s do the same for the origins
# Either use one single location, or a grid of locations, with a point for each cell of the population raster
loc = [Point(15.25, -4.24)]
loc_gdf = gpd.GeoDataFrame({'geometry':loc}, crs = {'init':'epsg:4326'}, geometry = 'geometry', index = [1])
origins_gdf = gpd.GeoDataFrame(res_df["geometry"], crs = epsg)
origins = gn.pandana_snap_c(G_time, origins_gdf, source_crs = epsg, target_crs = epsg_utm)
origins = [row.NN for idx,row in origins.iterrows()]
The Origin Destination matrix displays the time (in seconds) to reach the
OD = gn.calculate_OD(G_time, origins, destinations, fail_value = 9999999)
OD
array([[ 455.97075428, 384.80586467, 1584.46735752, ..., 750.99850665,
1479.42870708, 918.54000375],
[ 388.31285346, 302.93721397, 1659.05756661, ..., 762.84540117,
1531.00604159, 993.13021284],
[ 389.03973363, 436.96887641, 1669.84314267, ..., 678.33503068,
1446.49567111, 991.77735525],
...,
[1416.83559064, 1426.14071933, 1066.60141953, ..., 1034.53824266,
272.93524563, 840.09558931],
[1533.02969961, 1542.3348283 , 1182.7955285 , ..., 1150.73235163,
389.1293546 , 956.28969828],
[1533.02969961, 1542.3348283 , 1182.7955285 , ..., 1150.73235163,
389.1293546 , 956.28969828]])
OD = OD / 60
OD_df = pd.DataFrame(OD, columns = destinations, index = origins)
OD_df
| 7682 | 6020 | 900 | 7943 | 8072 | 9232 | 1692 | 3877 | 12841 | 2349 | ... | 3395 | 7112 | 3149 | 12123 | 3167 | 10595 | 8676 | 5101 | 12530 | 2806 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 4019 | 7.599513 | 6.413431 | 26.407789 | 10.780243 | 8.448830 | 19.711251 | 14.084332 | 23.769943 | 19.220016 | 26.969619 | ... | 20.444827 | 8.121287 | 25.594944 | 10.199661 | 19.642977 | 11.741221 | 23.444175 | 12.516642 | 24.657145 | 15.309000 |
| 687 | 6.471881 | 5.048954 | 27.650959 | 12.023413 | 9.692000 | 20.601789 | 13.958396 | 24.629566 | 19.638232 | 27.829241 | ... | 21.687997 | 6.146792 | 26.454566 | 8.317245 | 20.886147 | 10.393093 | 24.687345 | 12.714090 | 25.516767 | 16.552170 |
| 5402 | 6.483996 | 7.282815 | 27.830719 | 12.203172 | 9.871760 | 19.193283 | 12.549890 | 23.221060 | 18.229726 | 26.420735 | ... | 21.437101 | 4.738286 | 25.046060 | 6.908738 | 20.084285 | 8.984587 | 24.867105 | 11.305584 | 24.108261 | 16.529623 |
| 2685 | 3.311977 | 3.346300 | 22.120253 | 6.492707 | 4.161294 | 15.423714 | 9.796796 | 19.482407 | 14.932480 | 22.682083 | ... | 16.157291 | 5.240373 | 21.307408 | 5.912125 | 15.355441 | 7.453685 | 19.156639 | 8.229106 | 20.369609 | 11.021464 |
| 13068 | 4.446298 | 2.827633 | 24.002227 | 8.374680 | 6.043268 | 17.305688 | 11.678770 | 21.364381 | 16.814454 | 24.564056 | ... | 18.039265 | 6.273995 | 23.189382 | 7.163018 | 17.237414 | 8.957292 | 21.038612 | 10.111080 | 22.251583 | 12.903438 |
| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
| 1362 | 25.550495 | 25.705580 | 19.713259 | 19.408478 | 20.747305 | 10.756471 | 19.343176 | 11.315606 | 16.233197 | 8.501634 | ... | 12.104287 | 24.880330 | 4.478690 | 23.181197 | 14.112743 | 23.230521 | 22.206405 | 19.178873 | 6.485489 | 15.938162 |
| 8608 | 22.844689 | 22.999774 | 17.007453 | 16.702672 | 18.041499 | 8.050665 | 16.637370 | 8.609800 | 13.527391 | 5.795828 | ... | 9.398481 | 22.174524 | 1.772884 | 20.475392 | 11.406937 | 20.524715 | 19.500599 | 16.473067 | 3.779683 | 13.232356 |
| 9713 | 23.613927 | 23.769012 | 17.776690 | 17.471909 | 18.810737 | 8.819903 | 17.406607 | 9.379037 | 14.296629 | 6.565065 | ... | 10.167719 | 22.943761 | 2.542122 | 21.244629 | 12.176175 | 21.293952 | 20.269836 | 17.242304 | 4.548921 | 14.001593 |
| 1362 | 25.550495 | 25.705580 | 19.713259 | 19.408478 | 20.747305 | 10.756471 | 19.343176 | 11.315606 | 16.233197 | 8.501634 | ... | 12.104287 | 24.880330 | 4.478690 | 23.181197 | 14.112743 | 23.230521 | 22.206405 | 19.178873 | 6.485489 | 15.938162 |
| 1362 | 25.550495 | 25.705580 | 19.713259 | 19.408478 | 20.747305 | 10.756471 | 19.343176 | 11.315606 | 16.233197 | 8.501634 | ... | 12.104287 | 24.880330 | 4.478690 | 23.181197 | 14.112743 | 23.230521 | 22.206405 | 19.178873 | 6.485489 | 15.938162 |
125 rows × 24 columns
Testing the marketsheds
ma.generate_market_sheds(travel_surf, hf, out_file = os.path.join(out_path, 'marketsheds_test.tif'))
marketsheds = rio.open(os.path.join(out_path, 'marketsheds_test.tif'))
marketsheds.plot()
---------------------------------------------------------------------------
AttributeError Traceback (most recent call last)
Cell In[246], line 2
1 marketsheds = rio.open(os.path.join(out_path, 'marketsheds_test.tif'))
----> 2 marketsheds.plot()
AttributeError: 'DatasetReader' object has no attribute 'plot'